Crash course in Julia (programming)
Julia is a powerful/efficient/high-level computer programming language. You can get into interacting mode right-away with:
julia

Exit with exit(). One may also also use Wolfram's notebook concept:
jupyter-notebook
We will often need to install packages, which, back to julia, is done as follows:
using Pkg
Pkg.add("IJulia")
Then, from a terminal, jupyter-notebook opens a session in a browser, in which one can start a Julia session with a New invocation (and choosing julia there).
Let us now play with things computers are good at (e.g., with statistics). We'll need another package first:
import Pkg; Pkg.add("Distributions")
Once this is done (once for ever on a given machine), you can then be:
using Distributions
Let us generate ten thousands random points following a squared-uniform probability distribution, $X^2$.
lst=[rand()^2 for i=1:10^5]
and after
using Plots
histogram(lst)

The theoretical result is obtained by differentiating the cumulative function, $f_{X^2}=dF_{X^2}/dx$, with
$$F_{X^2}(x)\equiv\mathbb{P}(X^2\le x)$$
but $\mathbb{P}(X^2\le x)=\mathbb{P}(X\le\sqrt{x})=\sqrt{x}$, since the probability is uniform. Therefore:
$$f_{X^2}(x)={1\over2\sqrt{x}}$$
Let us check:
f(x)=1/(2*sqrt(x))
histogram(lst,norm=true)
plot!(f,.01,1, linewidth = 4, linecolor = :red, linealpha=0.75)
The ! means to plot on the existing display. This seems to work indeed:

Let's have a look at the distribution of inverse random numbers, also sampled from a uniform distribution between 0 and 1. This time, the support is infinite (it is $[1,\infty[$). The theoretical distribution is $f_{1/X}=d\mathbb{P}_{1/X}/dx$ with $\mathbb{P}_{1/X}=\mathbb{P}(1/X\le x)=\mathbb{P}(X\ge 1/x)=1-1/x$ (being uniform again), so that $f_{1/X}(x)=1/x^2$.
This time we'll use a lot of points to make sure we get the exact result:
@time lst = [1/rand() for i=1:10^8]
The @time allows us to benchmark the time and memory resources involved. As you can see, on a 2020 machine, it's pretty fast (for a hundred million points!)
0.427357 seconds (67.69 k allocations: 766.257 MiB, 0.65% gc time)
The result is also consequently very smooth:
@time histogram(lst,norm=true,bins=0:.01:10)
returning
4.644046 seconds (4.88 k allocations: 1.490 GiB, 1.74% gc time)
and, after also
f(x)=1/x^2
plot!(f,1,10,linewidth=4,linecolor=:red,linealpha=.75)

This is slightly off, clearly. The culprit is our distribution function, which density should be 1 at x=1 and is more than that. The problem is the norm option of histogram, that normalizes to what is plotted (or retained). And we have points beyond that. We can use this number to renormalize our theory plot. This is how many numbers we have:
sum(lst .< 10)
and this is the list of these numbers:
lst[lst .> 100]
When plotting pure functions, you might find that the density of points (PlotPoints in Mathematica) is not enough. This can be increased by specifying the step in the range, but in this case beware not to query your functions outside of its validity range, which would work otherwise. Compare:
plot(x->sqrt((15/x^2)-1),0,4,linewidth=5,linealpha=.5,linecolor=:red,ylims=(0,3))
plot!(x->sqrt((15/x^2)-1),0:.0001:3.8729,linewidth=2,linecolor=:blue,ylims=(0,3))

The first version can plot over the range 0–4 but does so by not plotting the full function, while the second (with x-steps of 0.0001 allowing for the nice curvature) shows everything but would return an error if going over the limit where the argument gets negative.
Let us now explore numerically a very interesting (and important) phenomenon. For that, we'll need the Cauchy function from the above-loaded "Distributions" package. This is a histogram of $10^5$ Cauchy-distributed points:
histogram(rand(Cauchy(),10^5))

Here the binning is mandatory:
histogram(rand(Cauchy(),10^5),bins=-10:.5:10,norm=true)

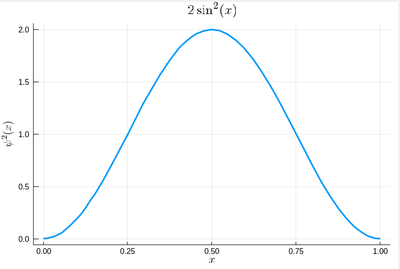
Let us now look at how to generate random numbers on our own. For instance, let us say we want to simulate the ground state of a particle in a box, with probability distribution over $[0,1]$ given by:
$$\psi^2(x)=2\sin^2(\pi x)$$
We can use unicode characters by entering \psi+TAB and call our density of probability ψ2, which in quantum mechanics is given by the modulus square of the probability amplitude (in 1D the wavefunction can always be taken real so we do not need worry about the modulus):
ψ2(x)=2*sin(pi*x)^2
That's our density of probability:
using LaTeXStrings
default(; w=3)
plot(ψ2,0,1, title=L"2\sin^2(x)", xlabel=L"x", ylabel=L"\psi^2(x)",dpi=150,legend=false)
where we decorated the plot with a lot of options, but the width which we gave as a default attribute, not to do it in the future (and since we like thick lines).
We will compute $\int_0^1\psi^2(x)\,dx$ by computing Riemann sums, i.e., as a sum of parallelograms defined in a partition of the support of the function (here $[0,1]$) cut into $n$ intervals. Calling $f$ the function to integrate, this is defined as:
$$R_n=\sum_{i=1}^{n}f(x_i)\delta(i)$$
a, b = 0, 1; # boundaries
lx=range(a, stop=b, length=11) # list of x intervals
δ=diff(lx) # widths of intervals
lxi=lx[1:end-1] # x_i points of parallelograms
We can now check the normalization:
sum([ψ2(lxi[i])*δ[i] for i=1:length(lxi)])
1.0
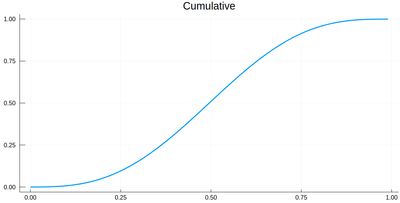
The cumulative is obtained as:
Ψ=[sum([ψ2(lxi[i])*δ[i] for i=1:j]) for j=1:length(lxi)]
This is the result for 100 intervals (length=101 above):
plot(lxi, Ψ, legend=false, title="Cumulative")
Now the generation of a random number following the initial distribution works as follows. We select randomly (uniformly) a number on the $y$-axis of the cumulative and find the corresponding $x$ such that $F(x)=y$. Those $x$ are $f$&nbps;distributed.
This can be simply achieved as follows (later we'll see a more sophisticated way):
yr=rand()
fx=lx[[findmin(abs.(Ψ.-rand()))[2] for i=1:10]]
Here are some positions where our ground state collapsed:
10-element Array{Float64,1}:
0.55
0.96
0.24
0.5
0.58
0.16
0.67
0.74
0.08
0.46
histogram(lx[[findmin(abs.(Ψ.-rand()))[2] for i=1:10^7]], bins=0:.01:1,norm=true)
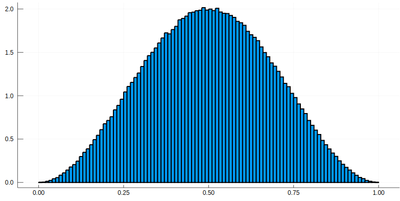
Note that the binning cannot be less than the $\delta$ step, otherwise we will get "holes" (and also the wrong normalization):
histogram(lx[[findmin(abs.(Ψ.-rand()))[2] for i=1:10^7]], bins=0:δ[1]/2:1,norm=true)
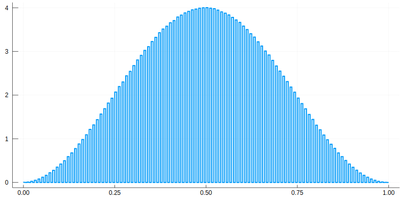
We would typically want to embed our results together in a function of our own:
function myRiemann(f, n)
a, b = 0, 1; # boundaries
lx=range(a, stop=b, length=n) # list of x intervals
δ=diff(lx); # widths of intervals
lxi=lx[1:end-1]; # x_i points of parallelograms
sum([f(lxi[i])*δ[i] for i=1:length(lxi)])
end
Then we can try to integrate all sorts of functions, e.g.,
function g(x)
exp(x)
end
myRiemann(g, 10^3)
The result is fairly close to the exact $e-1\approx1.718281828459045$:
1.717421971020861
However we'll find that our method is fairly limited:
scatter([abs(myRiemann(g, i)-(exp(1)-1)) for i=2:100:1002],
ylims=(10^-4,1), yscale=:log10, xticks=(1:11, 2:100:1002),
xlabel="Num steps", ylabel="error")
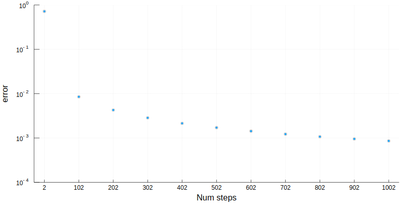
It is the purpose of numerical methods to learn how to use algorithms that are efficient, in the sense that they are accurate, fast and resource-effective.
It is easy to be brutal and terribly inefficient with a computer. In fact a fairly trivial enhancement of our method leads to considerable improvement:
function myRiemann(f, n, method="mid")
a, b = 0, 1; # boundaries
lx=range(a, stop=b, length=n) # list of x intervals
δ=diff(lx); # widths of intervals
if method == "left"
lxi=lx[1:end-1]; # x_i points of parallelograms
elseif method == "right"
lxi=lx[2:end];
else
lxi=(lx[1:end-1]+lx[2:end])/2;
end
sum([f(lxi[i])*δ[i] for i=1:length(lxi)])
end
Of course, other people already wrote such functions, that are available through packages. For numerical integration of 1D functions, one can use QuadGK which is based on the so-called Gauss–Kronrod quadrature formula, which basically figures out how to best choose the points $x_i$ and how to weight them:
using QuadGK
@time quadgk(g, 0, 1)
Unlike our naive implementation, the result is pretty accurate:
0.000034 seconds (9 allocations: 304 bytes) (1.718281828459045, 0.0)
That's the best we can achieve (with mid-points and a hundred-million intervals!)
@time myRiemann(g, 10^8)
9.762264 seconds (500.00 M allocations: 8.941 GiB, 11.01% gc time) 1.7182818284590453
It differs from the exact result only in the last digit, but took about 10s & 9GiB of memory.
So much for integration. How about the reverse process, i.e., derivation. We can use the diff command that computes finite differences:
diff(1:.5:10)
returns an array of 18 elements, all equal to 0.5. In this way, it is easy to compute a finite-difference derivative:
plot(diff([sin(pi*x) for x=0:.01:1])./.01)
This is a cosine. Importantly for numerical methods, one should look for simple tricks that maximise numerical accuracies. The idea behind differentiation comes from Taylor expansion:
\begin{equation} \label{eq:taylorf} f(x\pm\epsilon)=f(x)\pm\epsilon f'(x)+{\epsilon^2\over2} f(x)\pm{\epsilon^3\over6} f'(x)+o(\epsilon^4) \end{equation}
that we wrote for both $\pm\epsilon$, and using the $o$-notation to mean that terms we neglected are of the order of $\epsilon^4$. From this, one can get an obvious numerical approximation for the derivative
$$f'(x)\approx{f(x\pm\epsilon)-f(x)\over\epsilon}+o(\epsilon)$$
meaning we can expect an error of the order of $\epsilon$ with this method. Observe however how one can sneakily cancel the $\epsilon^2$ term from \eqref{eq:taylorf} by subtracting $f(x-\epsilon)$ from $f(x+\epsilon)$:
$$f(x+\epsilon)-f(x-\epsilon)=2\epsilon f'(x)+{\epsilon^3\over 3}{f(x)}+o(\epsilon^4)$$
so that we have a much better approximation by computing with this slight variation:
$$f'(x)={f(x+\epsilon)-f(x-\epsilon)\over 2\epsilon}+o(\epsilon^2)$$
since $\epsilon^2\ll\epsilon$. The computational cost is the same, the numerical gain is considerable. Let's write a function. We'll it call Dl for "Differentiate list":
function Dl(f, δx, method="central")
if method=="left"
diff(f)/δx
elseif method=="central"
[f[i+1]-f[i-1] for i=2:length(f)-1]/(2*δx)
end
end
Let us compare the two methods for
δx=0.1;
mysin=[sin(pi*x) for x in 0:δx:1]
We find fairly identical-looking results:
plot([Dl(mysin,δx,"central"), Dl(mysin,δx,"left")], legend=false)

This error is actually quite large if looked at properly: (note the comparison to lists of different size, the central method removes two points)
plot([
Dl(mysin,δx,"central")-[pi*cos(pi*i) for i in δx:δx:1-δx],
Dl(mysin,δx,"left")-[pi*cos(pi*i) for i in 0:δx:1-δx]
], legendtitle="Method", legend=:bottomleft, label=["central" "left"])

This idea (and method) can be further exploited. By working out how to cancel the next-order terms, we arrive at the so-called Five-point stencil:
$$f'(x) \approx \frac{-f(x+2\epsilon)+8 f(x+\epsilon)-8 f(x-\epsilon)+f(x-2\epsilon)}{12\epsilon}$$
which we can add to our function:
function Dl(f, δx, method="central")
if method=="left"
diff(f)/δx
elseif method=="central"
[f[i+1]-f[i-1] for i=2:length(f)-1]/(2*δx)
elseif method=="fivepoints"
[-f[i+2]+8f[i+1]-8f[i-1]+f[i-2] for i=3:length(f)-2]/(12*δx)
end
end
Comparing their errors:
plot([
Dl(mysin,δx,"fivepoints")-[pi*cos(pi*i) for i in 2δx:δx:1-2δx],
Dl(mysin,δx,"central")-[pi*cos(pi*i) for i in δx:δx:1-δx],
Dl(mysin,δx,"left")-[pi*cos(pi*i) for i in 0:δx:1-δx]
], legendtitle="Method", legend=:bottom,
label=["5 points" "central" "left"], ylims=(-.005,.001))

If we want a more quantitative study at the errors, we can look at how it scales for various $\epsilon$. This will allow us to look at logarithmic steps:
lδx=[10.0^(-k) for k in 1:7]
err1=[mean(abs.(Dl([sin(pi*x) for x in 0:δx:1], δx, "left")-[pi*cos(pi*x) for x in 0:δx:1-δx])) for δx in lδx]
err2=[mean(abs.(Dl([sin(pi*x) for x in 0:δx:1], δx, "central")-[pi*cos(pi*x) for x in δx:δx:1-δx])) for δx in lδx]
err3=[mean(abs.(Dl([sin(pi*x) for x in 0:δx:1], δx, "fivepoints")-[pi*cos(pi*x) for x in 2δx:δx:1-2δx])) for δx in lδx]
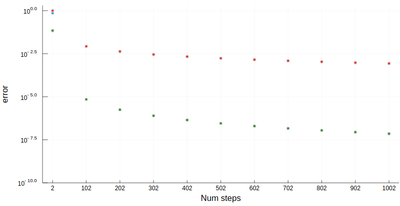
plot([abs.(err1) abs.(err2) abs.(err3)],
yscale=:log10, label=["left" "central" "5 points"],
xlabel="10^-k step", ylabel="Mean abs error", lc=[:red :blue :green])
plot!([x->10.0^(-x) x->10.0^(-2x) x->10.0^(-4x)],1,7,
yscale=:log10, ylims=(10^(-15),1), lc=[:red :blue :green], ls=:dash, label=["" "" ""])
with the beautiful result:

As one can see, the best method worsens for $\epsilon<10^{-4}$ so it will actually work better (and faster) for not too small an $\epsilon$.
Let us now turn to another typical numerical method: a root-finding algorithm. We used a so-called bisection method previously when generating our own random numbers, which is easy to implement but not very efficient. A simple but good improvement is the Newton–Raphson method (Newton used it for the particular case of polynomials; Raphson extended it to arbitrary functions). The idea is to start with a blind guess $x_0$ around the solution and replace the real function $f(x)$ by a linear approximation with the same slope at $x_0$, i.e., $h(x)=f'(x_0)x+\beta$ with $\beta$ such that $h(x_0)=f(x_0)$ which yields
$$h(x)=f'(x_0)[x-x_0]+f(x_0)$$
Now a better guess for the zero is found by solving $h(x)=0$ which has solution $x=x_0-(f(x_0)/f'(x_0))$. Of course the procedure can be iterated so the algorithm gives us the always better approximations:
$$x_{n+1}=x_n-{f(x_n)\over f'(x_n)}\,.$$
Let us find the zeros of $f(x)=\sin(x^2)$ between 0 and $\pi$. It's good to know beforehand the sort of results we expect (we added vertical lines by eye):
psin2=plot(x->sin(x^2), 0, π,
ylabel=L"\sin(x^2)",xlabel=L"x",lw=3,
legend=false)
vline!([0, 1.77, 2.51, 3.07], line = (:red, :dash, 1))
hline!([0], line = (:black, :dot, .75))

The formula gives us:
$$x_{n+1}=x_n-{\sin x_n^2\over 2x_n\cos x_n^2}$$
We can write a function that takes $f/f'$ as an input (that we have to compute ourselves) that computes such iterations until the procedure differs by less than some $\epsilon$:
function NewtonRaphson(f, x0)
ϵ=10^-16
x1=x0+10ϵ
while abs(x1-x0)>ϵ
x0=x1
x1=x0-f(x0)
end
x1
end
This provides the values found in the $[0,\pi]$ interval:
lsols=[NewtonRaphson(x->sin(x^2)/(2*x*cos(x^2)), x) for x in 0:.01:π]
We can plot the solutions together with where was the starting point:
psin2=plot(x->sin(x^2), 0, π,
label=L"\sin(x^2)",xlabel=L"x",
legend=:bottomleft)

We found many, in particular some outside the $[0,\pi]$ range (this was the case when the tangent was sending us far away). We can check these are indeed all solutions:
plot(abs.(sin.(lsols.^2)), yscale=:log10, ylims=(10^-40,10^-10), legend=false)

To actually extract the values, we could use unique(), but this would return a lot of different values for the 0 zero, so we'd round this:
lsolss=sort(unique(round.(lsols,digits=10)))
These are the said solution in the sought range:
lsolss[findall(x-> 0<=x<=π, lsolss)]
5-element Array{Float64,1}:
-0.0
0.0
1.7724538509
2.5066282746
3.0699801238
-0.0 and 0.0 are seen as different in julia, surely something that'll be fixed eventually. We have four solutions, as is clear from the initial graph.
There is a Roots package that has a function fzero() that achieves something similar:
fzero(x->sin(x^2), 2)
1.772453850905516
Let us now turn to a typical numerical problem, solving differential equations. We'll start with Ordinary Differential Equations (ODE). Namely, we consider equations of the type:
$$\label{eq:ODE1} y'(t)=f(t,y(t))$$
The most famous algorithm is Euler's method and the most popular the RK4 variant of the Runge–Kutta methods. In between are a variety of possible implementations, many related in some way or the other. Methods can be explicit (the solution only depends on quantities already known) or implicit (the solution involves other unknowns which requires solving intermediate equations). The backward Euler method is an implicit method. Euler's method is the simplest one and the typical starting point to introduce the basic idea. Not surprisingly, it is also one of the least efficient. We discretize time into steps $t_1$, $t_2$, $\cdots$, $t_n$ separated by the same interval $h$. The solutions at these times are denoted $y_n=y(t_n)$. Back to \eqref{eq:ODE1}, approximating $y'(t)\approx (y(t+h)-y(t))/h$, we have:
$$y_{n+1}=y_n+hf(t_n,y_n)$$
This is very simple since from the solution $y_n$ at time $t_n$ we can compute the solution at $t_{n+1}$. Explicitly.
Let us solve, for instance, $y'=-y$, whose solution is the exponential function.
npts = 100; # number of points
y=0.0*collect(1:npts); # predefine the array
h=.1; # timesteps, so total time = npts*h=10
# The differential equation to solve
function f(t,y)
-y
end
y[1]=1; # initial condition
# The numerical method (Euler's)
for i=1:npts-1
y[i+1]=y[i]+h*f((i-1)*h,y[i])
end
And we can now compare the numerical solution to the real one:
plot(((1:npts).-1)*h, [y[i] for i=1:npts], label="Euler", xlabel="t", ylabel="y'=-y")
plot!(x->exp(-x),0, 10, label="Exact")

Let us try the more difficult example:
$$y'={y\over 2}+2\sin(3t)\label{eq:PaulODE}$$
whose solution can be found with an integrating factor [1]
$$y=-{24\over37}\cos(3t)-{4\over37}\sin(3t)+\left(y(0)+{24\over 37}\right)e^{t/2}$$
In this case the differential equation is defined as:
function f(t,y)
(y/2)+2sin(3t)
end
And the method can be quickly implemented as:
tmax=4*pi;
h=0.0001;
npts=round(Int64,tmax/h);
y=0.0*collect(1:npts);
y[1]=-24/37
for i=1:npts-1
ti=(i-1)*h
y[i+1]=y[i]+h*f(ti,y[i])
end
with various repetitions of
plot!(((1:npts).-1).*h, y,
xlabel="t", ylabel="y", label="h=$(h)",
legend=:bottomleft,ylims=(-2,.75),lw=3)
to see the effect of different time steps:

Comparing to the exact solution shows that the error gets quickly sizable, even with a lot of points:
plot(abs.([-(24/37)*cos(3t)-(4/37)*sin(3t) for t in ((1:npts).-1).*h]-y),
lw=3, color=:red, xlabel="index array", ylabel="error", legend=false)

We can analyze the error in more details:
hlist=[10.0^(-k) for k=1:6];
herror=[];
for h in hlist
npts=round(Int64,tmax/h);
y=0.0*collect(1:npts);
y[1]=-24/37;
@time for i=1:npts-1
ti=(i-1)*h
y[i+1]=y[i]+h*f(ti,y[i])
end
append!(herror,mean(abs.([-(24/37)*cos(3t)-(4/37)*sin(3t) for t in ((1:npts).-1).*h]-y)))
end
produces on wulfrun2:
0.000689 seconds (880 allocations: 15.813 KiB) 0.003283 seconds (11.78 k allocations: 203.766 KiB) 0.025673 seconds (136.18 k allocations: 2.270 MiB) 0.314037 seconds (1.38 M allocations: 22.979 MiB, 1.45% gc time) 2.519780 seconds (13.82 M allocations: 230.066 MiB, 1.00% gc time) 26.261842 seconds (138.23 M allocations: 2.247 GiB, 2.43% gc time)
and a nice linear decay of the error with the timestep (so no-round off error, because essentially we cannot take a smaller timestep):

Euler's method works but it is not very accurate. It can be improved by considering a better estimation for the tangent. The idea is to use not the tangent of $y$ at $t_n$ only but also at $t_{n+1}$ and take the average:
$$y_{n+1}=y_n+{h\over 2}[f(t_n,y_n)+f(t_{n+1},y_{n+1})]$$
Of course, we do not know $y_{n+1}$, so the method would appear to become implicit, and we will deal with such a case later. But here, simply, we merely use Euler's method to estimate it:
$$y_{n+1}=y_n+hf(t_n,y_n)$$
so that we have an explicit formula again:
$$y_{n+1} = y_n + {h\over2} [f(t_n, y_n) + f(t_n + h, y_n + h f(t_n, y_n))]$$
This is known as Heun's method. It is one of the simplest of a class of methods called predictor-corrector algorithms (Euler's method is only a predictor method). In code:
@time for n=1:npts-1
tn=(n-1)*h-t0;
y[n+1]=y[n]+(h/2)*(f(tn,y[n])+f(tn+h,y[n]+h*f(tn,y[n])))
end
We can similarly compute the mean absolute error with this method:
herror=[];
for h in hlist
npts=round(Int64,tmax/h);
y=0.0*collect(1:npts);
y[1]=-24/37;
@time for n=1:npts-1
tn=(n-1)*h;
y[n+1]=y[n]+(h/2)*(f(tn,y[n])+f(tn+h,y[n]+h*f(tn,y[n])))
end
append!(herror,mean(abs.([-(24/37)*cos(3t)-(4/37)*sin(3t) for t in ((1:npts).-1).*h]-y)))
end
still on wulfrun2:
0.000713 seconds (2.00 k allocations: 33.391 KiB) 0.009036 seconds (23.08 k allocations: 380.391 KiB) 0.051665 seconds (249.26 k allocations: 3.995 MiB) 0.385209 seconds (2.51 M allocations: 40.236 MiB, 2.70% gc time) 3.921418 seconds (25.13 M allocations: 402.639 MiB, 1.38% gc time) 39.136207 seconds (251.33 M allocations: 3.932 GiB, 1.99% gc time)
This is more resource intensive but now we get a much better $h^2$ error:
scatter(herror, xlabel="time step", ylabel="mean abs error",
xticks=(1:6, string.(hlist)), legend=false, yscale=:log10)
plot!(x->100*herror[1]*10.0^-(2x), 1, 6, ls=:dot, lw=3)

We now cover one of the most powerful predictor-corrector algorithms of all,
one which is so accurate that most computer algorithms use it as default: the fourth-order Runge-Kutta Method. RK methods are optimized predictor-corrector methods, so they push the idea of Heun's method to the top. The RK4 method finds the optimum correction-prediction (in a way which we do not detail here) to involve these intermediate points:
\begin{align}
k_1 &= h\ f(t_n, y_n)\,, \\
k_2 &= h\ f\left(t_n + \frac{h}{2}, y_n + \frac{k_1}{2}\right)\,, \\
k_3 &= h\ f\left(t_n + \frac{h}{2}, y_n + \frac{k_2}{2}\right)\,, \\
k_4 &= h\ f\left(t_n + h, y_n + k_3\right)\,,
\end{align}
and then the calculation is explicit:
\begin{align} t_{n+1} &= t_n + h\,, \\ y_{n+1} &= y_n + \tfrac{1}{6}\left(k_1 + 2k_2 + 2k_3 + k_4 \right)\,. \end{align}
In code:
@time for n=1:npts-1
tn=(n-1)*h-t0;
k1=h*f(tn,y[n]);
k2=h*f(tn+(h/2),y[n]+k1/2);
k3=h*f(tn+(h/2),y[n]+k2/2);
k4=h*f(tn+h,y[n]+k3);
y[n+1]=y[n]+(1/6)*(k1+2*k2+2*k3+k4)
end
The error analysis on the same problems as above now yields a very different result:
herror=[];
for h in hlist
npts=round(Int64,tmax/h);
y=0.0*collect(1:npts);
y[1]=-24/37;
@time for n=1:npts-1
tn=(n-1)*h;
k1=h*f(tn,y[n]);
k2=h*f(tn+(h/2),y[n]+k1/2);
k3=h*f(tn+(h/2),y[n]+k2/2);
k4=h*f(tn+h,y[n]+k3);
y[n+1]=y[n]+(1/6)*(k1+2*k2+2*k3+k4)
end
append!(herror,mean(abs.([-(24/37)*cos(3t)-(4/37)*sin(3t) for t in ((1:npts).-1).*h]-y)))
end
0.006093 seconds (3.91 k allocations: 64.688 KiB) 0.007069 seconds (41.92 k allocations: 674.766 KiB) 0.071775 seconds (437.74 k allocations: 6.871 MiB) 0.805397 seconds (4.40 M allocations: 68.998 MiB, 1.76% gc time) 6.734973 seconds (43.98 M allocations: 690.260 MiB, 1.41% gc time) 77.495034 seconds (439.82 M allocations: 6.741 GiB, 1.57% gc time)
scatter(herror, xlabel="time step", ylabel="mean abs error",
xticks=(1:6, string.(hlist)), legend=false, yscale=:log10)
plot!(x->10^4*herror[1]*10.0^-(4x), 1, 6, ls=:dot, lw=3, ylims=(10^-15,10^-3))

The gain from $h=0.001$ to $h=0.0001$ is marginally better, and for smaller still $h$, it actually becomes worse, due to rounding errors. It works so well that many, if not most, resources would use the RK4 to integrate ODE.
We now turn to the case of coupled equations, in which case the method reads
$$\vec y'=\vec f(t,\vec y)$$
where
$$ \vec y\equiv \begin{pmatrix}
y_1\\y_2
\end{pmatrix} \quad\text{and}\quad \vec f\equiv \begin{pmatrix}
f_1\\f_2
\end{pmatrix}\,. $$
The methods are the same as before just applying directly to the vectors instead. An interesting coupled system of ODE, namely, the Volterra-Lotka model of prey-predator interactions. This is modelled with the pair of coupled ODE:
\begin{align}
\frac{dx}{dt} &= \alpha x - \beta x y, \\
\frac{dy}{dt} &= \delta x y - \gamma y,
\end{align}
The functions are implemented as:
function f1(t,y1, y2)
α*y1-β*y1*y2
end
function f2(t,y1, y2)
δ*y1*y2-γ*y2
end
Some common initialization for all methods, in particular, we now need two arrays of values to store the evolution of $x$ (in y1) and $y$ (in y2):
tmax=100
h=.001
npts=round(Int,tmax/h)
y1=0.0*collect(1:npts);
y2=0.0*collect(1:npts);
y1[1]=1;
y2[1]=.1;
α=2/3;
β=4/3;
γ=1;
δ=1;
Euler's method in this case reads:
$$\vec y_{n+1}=\vec y_n+h\vec f(t_n,\vec y_n)$$
or, breaking down this equation componentwise:
$$\begin{align} y_{1,n+1}&=y_{1,n}+hf_1(t_n,y_{1,n},y_{2,n})\\ y_{2,n+1}&=y_{2,n}+hf_2(t_n,y_{1,n},y_{2,n}) \end{align}$$
In code:
@time for i=1:npts-1
y1[i+1]=y1[i]+h*f1((i-1)*h,y1[i],y2[i])
y2[i+1]=y2[i]+h*f2((i-1)*h,y1[i],y2[i])
end
Heun's version reads:
$$\vec y_{n+1}=\vec y_n+{h\over 2}\left(\vec f(t_n,\vec y_n)+\vec f(t_{n+1},\vec y_n+h\vec f(t_i,\vec y_n)\right)$$
or, breaking down this equation componentwise:
$$\begin{align}
y_{1,n+1} &= y_{1,n} + {h\over2} [f(t_n, y_{1,n}, y_{2,n}) + f(t_n + h, y_{1,n} + h f(t_n, y_{1,n}, y_{2,n}), y_{2,n})]\\
y_{2,n+1} &= y_{2,n} + {h\over2} [f(t_n, y_{1,n}, y_{2,n}) + f(t_n + h, y_{1,n}, y_{2,n} + h f(t_n, y_{1,n}, y_{2,n}))]
\end{align}$$
In code; where this time we'll find both more convenient and efficient to define auxiliary quantities, that are furthermore invoked more than once:
@time for n=1:npts-1
tn=(n-1)*h;
f1left=f1(tn,yH1[n],yH2[n])
f2left=f2(tn,yH1[n],yH2[n])
f1right=f1(tn+h,yH1[n]+h*f1left,yH2[n]+h*f2left)
f2right=f2(tn+h,yH1[n]+h*f1left,yH2[n]+h*f2left)
yH1[n+1]=yH1[n]+(h/2)*(f1left+f1right)
yH2[n+1]=yH2[n]+(h/2)*(f2left+f2right)
end
RK4's version reads:
$$ \begin{align} t_{n+1} &= t_n + h\,, \\ \vec y_{n+1} &= \vec y_n + \tfrac{1}{6}\left(\vec k_1 + 2\vec k_2 + 2\vec k_3 + \vec k_4 \right)\,. \end{align} $$
in terms of the intermediate quantities:
$$ \begin{align} \vec k_1 &= h\vec f(t_n, \vec y_n)\,, \\ \vec k_2 &= h\vec f\left(t_n + \frac{h}{2}, \vec y_n + \frac{\vec k_1}{2}\right)\,, \\ \vec k_3 &= h\vec f\left(t_n + \frac{h}{2}, \vec y_n + \frac{\vec k_2}{2}\right)\,, \\ \vec k_4 &= h\vec f\left(t_n + h, \vec y_n + \vec k_3\right)\,, \end{align} $$
In code
@time for n=1:npts-1
tn=(n-1)*h;
# Intermediate steps
k11=h*f1(tn,yRK1[n],yRK2[n]);
k12=h*f2(tn,yRK1[n],yRK2[n]);
k21=h*f1(tn+(h/2),yRK1[n]+k11/2,yRK2[n]+k12/2);
k22=h*f2(tn+(h/2),yRK1[n]+k11/2,yRK2[n]+k12/2);
k31=h*f1(tn+(h/2),yRK1[n]+k21/2,yRK2[n]+k22/2);
k32=h*f2(tn+(h/2),yRK1[n]+k21/2,yRK2[n]+k22/2);
k41=h*f1(tn+h,yRK1[n]+k31,yRK2[n]+k32);
k42=h*f2(tn+h,yRK1[n]+k31,yRK2[n]+k32);
# Real step
yRK1[n+1]=yRK1[n]+(1/6)*(k11+2*k21+2*k31+k41);
yRK2[n+1]=yRK2[n]+(1/6)*(k12+2*k22+2*k32+k42);
end
Results can be shown in two ways:
1 - As the respective solutions. This shows the oscillations of predators and preys
pRK=plot([[yRK1[i] for i=1:240], [yRK2[i] for i=1:240]],
lw=3, xlabel="time", ylabel="populations", label="RK4")

2 - As a phase-space solution, showing the simultaneous state of the system (here, ecosystem), with time as a parametric parameter:
psRK=plot([(yRK1[i], yRK2[i]) for i=1:npts],
lw=1, xlabel="x", ylabel="y", legend=false)

This comparing the three methods together (the various plots have been evaluated as pE, pH, etc.):
plot(pE,pH,pRK,psE,psH,psRK,layout=(2,3),dpi=150)

The time-evolutions at the top are over considerably fewer cycles for Heun & RK4 methods than those simulated to produce the phase space solutions, bottom. You can see how the RK4 method stays stable (which is the correct behaviour) while the Heun method slowly drifts towards the stable point.